How human evolution has been speeding up
Providing some of the missing background for new research on natural selection from ancient DNA
Nineteen years ago, I wrote about my collaborative work showing that human evolution has recently accelerated. Our study found parts of the genome where new variants had arisen within the last 40,000 years and then grown rapidly in frequency. Beyond a certain point, that pattern should be very unusual unless the new variant helped its carriers to survive or reproduce. This is positive natural selection. In the samples we studied, from European, African, and East Asian ancestry groups, we found hundreds of apparent examples.
We saw right away that hundreds of adaptive variants in the last 40,000 years implied a rate of change an order of magnitude faster than seemed to be true across our longer-term evolutionary history. Moreover, if positive selection had been this pervasive across a longer time—as much as 100,000 years—then hundreds of new mutations should have swept to fixation in that time. Those would have left wide wakes of surrounding genome depauperate of variation. Even nineteen years ago we knew such regions were very rare.
We had to conclude that the last 40,000 years were not typical of the longer span of human evolution. Somehow positive selection had sped up.
But why? For one thing, human environments have been changing in unprecedented ways. Agriculture and animal domestication brought new foods, new diseases, new modes of social organization. Long before that, modern humans who moved into new regions grappled with new regimes in sunlight, day length, altitude, and aridity. Many of the new genes under selection—affecting traits like pigmentation, metabolism, milk digestion, kidney function, and immunity—helped to adapt us to these new environments.
For another thing, human populations had grown in numbers by more than a hundredfold. From the thin populations of hunter-gatherers to urban centers and rural landscapes covered in villages and farms, the most pervasive fact about humanity was this increase in numbers. Useful mutations are incredibly rare. One hundred times more people is like buying a hundred times more lottery tickets.
We put together these two ideas into a theory unifying the observations from archaeology and anthropology with the observations from human genetics. Our model helped explain why evidence for positive selection was common not just in populations with a legacy of early agriculture but also in Africa where the resource intensification of the Late Stone Age involved the growth and connectivity of hunter-gatherer groups.
I’m recounting that story because last week, almost twenty years later, the recent, rapid evolution of our species was again in the news. Nature has published work by Ali Akbari and coauthors from David Reich’s research group, based on the largest-ever sample of more than 15,000 genomes of ancient individuals. They found statistical support for more than 400 genetic variants subject to positive natural selection in western Eurasia during the last 10,000 years.
Like us, Akbari and coworkers interpret this as an acceleration. I’m really pleased that this line of evidence from ancient DNA confirms our model.
Akbari and collaborators released a preprint of their work two years ago, and I included their work in my top 10 list of papers for 2024. Since then, their team and others have followed up with additional work. Recently this group in a preprint led by Alison Barton showed that ancient genomes from East Asia reflect a similar pattern of evolution.
I have a lot to share about this growing body of work on selection from ancient DNA and how it connects to my own. These ancient DNA results are strongly aligned with a broad foundation of previous research. In this post I’ll briefly reflect on my personal story studying selection and the breadth of work in this area from the early 2000s. This history shows how ancient DNA can illuminate some aspects of recent evolution that are challenging from retrospective studies from living people’s genomes.
What ancient genomes add
Akbari and coauthors write that when we study selection in samples of living people, it is like “being trapped in the present and studying the scars left by selection on the genomes of descendants”.
I understand this feeling. I was trained as a paleoanthropologist, so my first thought is always of samples from skeletons. I learned population genetics from Frank Livingstone, whose research on the genetic basis of malaria resistance taught me a great deal about how a new pathogen could create the circumstances for rapid selection on dozens of different genes. My background in studying Neanderthals and later people made it obvious that the skeletons of early farmers had evolved really fast in comparison to the much slower evolution of humans in the Pleistocene.
One day in my graduate training, I walked into the office of the archaeologist Henry Wright, known for his work on early state formation. “Agriculture should have sped up human evolution!” I said. I explained how the changes in disease ecology, diet, and life history would create the circumstances for rapid selection.
In retrospect, I doubt the idea was very new to Henry. In any event, I remember him taking a beat before replying, “Well, yes. That makes sense. But how can you show it?”
So I was paying close attention in the early 2000s when genetic evidence for recent positive selection began to build. Pardis Sabeti from Eric Lander’s research group led the development of a test for selection based on long-range linkage across a portion of a chromosome. They highlighted a gene known to be associated with malaria resistance in Mediterranean and North African groups, G6PD, as well as an immune system gene, CD40. In a later paper, that group characterized the haplotype structure around the gene for lactase, LCT. Strong recent selection on many other genes was characterized in that decade—genes related to pigmentation, diet, immunity, and classical anthropological traits like earwax consistency and hair thickness.
Before long, several research teams went on hunts for selection across the whole genome. Benjamin Voight from Jonathan Pritchard’s research team led a study of the haplotype structure of genomes from the International HapMap samples. They found more than 600 regions with statistical evidence of long, common haplotypes consistent with selection in the three regional samples. My collaborators Eric Wang and Robert Moyzis developed a related approach to examine a sample of genomes developed by Perlegen Sciences, and my collaboration with them in 2007 applied that method to the HapMap samples also. Analysis of samples from more than fifty populations that were included within the Human Genome Diversity Project by Pritchard’s group, in work led by Joe Pickrell, leveraged between-population methods to replicate some of the earlier studies of other samples while adding more loci and populations. All this work showed that positive selection had been very widespread in the recent past.
These approaches had many limitations. They worked on new mutations that were strongly selected. New malaria resistance variants like G6PD deficiency and the Duffy null blood type stood out, while the adaptive effects on malaria of standing variants like O blood type did not.
Other patterns of selection were likewise invisible. Some classic samples of natural selection in humans, like sickle cell or human leukocyte antigen (HLA) alleles, were cases of balancing selection. Some of these—especially the malaria adaptations—were new variants that appeared within the past 10,000 years. But their low frequencies or complicated haplotype structure meant the early whole-genome samples had little statistical power to detect them.
As the data grew, it became clear that selection on phenotypes often affected both new mutations and standing variants. For example, pigmentation is an oligogenic trait, influenced strongly by around 25 genes, some of which have new selected mutations and others show directional selection on standing variants. Even lactase persistence, famous for new strongly-selected mutations in several populations, also has seen standing variants selected in other populations.
Being trapped in the present imposes another limitation. Human prehistory was far from static. Some groups grew in numbers and in geographic range, mixing in the process. Others declined or became extinct. Our genealogies all thread through this complicated history, tangled up in some places and times much more than others. The statistical models for identifying variants under selection often assumed a static population, and that meant a better understanding of population history was badly needed.
To understand the details of both selection and population history, looking beyond present-day samples back at skeletal remains is essential. I’ve always seen the complexity of population history as more of an opportunity than impediment, because population history and selection are not isolated from each other; they interact.
When I started this work twenty years ago, it was already clear that early farmers had brought Neolithic lifestyles into Europe from southwest Asia, and Bronze Age Europe had an influx of steppe peoples. I traveled to many collections with Bronze Age and Neolithic skeletal remains to try to lay a groundwork for understanding how selection and demographic histories had interacted. Despite some very positive reviews, funding to expand my pilot project never worked out. I still think there’s a great deal of promise in such studies, and I hope to return to them. Skeletons combine signals of genetic relationship with variation from environment and life history.
Ancient DNA from the skeletons is more direct. Beginning in the early 2010s, researchers began to work on a small scale with samples from medieval, Bronze Age, and Neolithic contexts from Europe. They often examined genetic variants known to be under recent selection, especially around the lactase gene, but also variants related to pigmentation. The lower frequency of selected variants in these ancient samples confirmed some of the retrospective findings from present-day samples.
Sometimes a recently-selected variant was more common than expected in an ancient group; other times it only showed up in samples long after it supposedly arose. In a 2015 paper, Aaron Sams—my former PhD student—described how ancient DNA was a valuable addition to allele age estimation, helping refine the timeline of selection.
By that year, a few hundred ancient genomes of modern humans enabled David Reich’s research group to carry out a genome-wide screen for selection. They characterized a dozen selected variants in that study, led by Iain Mathieson. Mathieson has subsequently led a variety of studies on how DNA evidence can shed light on the phenotypes of ancient people.
Strengths of the new work
A big weakness of ancient DNA work across the last decade has been the small size of samples. The new work has more than 15,000 genomes, and that helps a lot. To see how much, it’s useful to compare Akbari and coworkers’ results with the new preprint from Alison Barton and coworkers, in which the same team applies the same statistical method to 1800 genomes from eastern Eurasia. In that preprint, they identify 40 variants under selection. The larger study by Akbari and coworkers identified more than 400.
The details of the statistical approach helped to further increase the signal. They applied a generalized linear mixed model, in which they evaluated the linear fit of each variant’s frequency with time, controlling for the network across all genomes. The effect is to leverage the variation in space and time of all the samples as a control on demographic history. It’s a little bit like finding asteroids by finding bright spots that move against the background field of stars—except the stars are jittering around and some move quite a bit.
Still, the study has some weaknesses that must be kept in mind. Every study of natural selection in DNA works with mathematical models of how natural selection has been patterned. A test based on one model will miss other patterns of selection. The model developed by Akbari and coworkers assumes a constant selection coefficient, and if substantial changes in the strength or direction of selection happened, those cases were likely missed.
The time series of ancient genomes reveals not only mutations that grew after arising recently but also changes to the frequencies of standing variants that have existed in the population for much longer. Akbari and coauthors estimate that around a third of their loci were new mutations, with selection on standing variants making up a larger fraction of their observations.
Earlier work was generally limited to the strongest cases of selection. Here, Akbari and coworkers have highlighted some cases where the frequency of a selected variant increased by as little as four or five percent. This helps enormously in understanding selection on traits that are influenced by multiple genes.
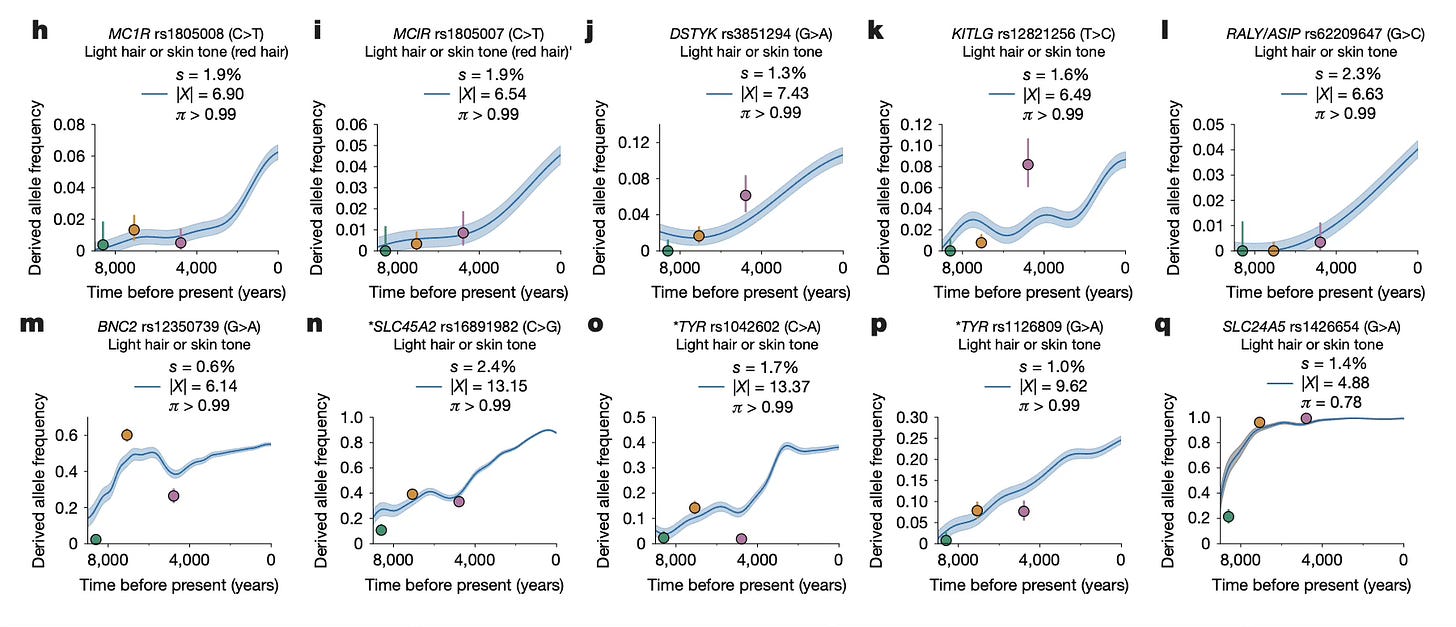
Pigmentation-related genes are a very clear illustration. It’s far from new to observe that lighter pigmentation was directionally selected in Europe and nearby parts of Asia for the last 10,000 years. Variants related to pigmentation involving genes like SLC24A5 and OCA2/HERC2 were among the earliest examples of recent strong selection to be characterized in the early 2000s. The network of protein-coding genes involved in pigmentation phenotypes is pretty well understood, and both coding and noncoding variants that affect those phenotypes are known from many populations. Akbari and coworkers were able to reproduce results on genes previously found to be under strong selection in Europe, and further showed recent selection on standing variants and more weakly-selected variants.
Most phenotypes are polygenic, with a large number of variants influencing their variation. When such phenotypes come under directional selection, many variants may change in frequency to small degree.
This leads to the most controversial aspect of the new study. Akbari and coworkers looked at the evolution of phenotypes through means of polygenic risk scores (PRS). In brief, a PRS is a statistical combination of genotypes across many loci that have been shown to be associated with a trait. Those associations are usually estimated by measuring phenotypes and genotyping tens of thousands of living people in large-scale samples such as the UK Biobank. The idea is that if you can find dozens of polymorphisms associated with a trait, you can then combine them into a single number. This score expresses an individual’s probability of developing the trait, given their genotypes.
The PRS approach has many known limitations. Since nearly all traits are influenced by environments in addition to genes, PRS scores are not usually very accurate for any given individual. Their usefulness is more in identifying groups within the population that may have higher risk of a disease, for example, or lower response to a medication.
When studying ancient genomes, there are additional challenges. Genetic associations estimated in one population are based on the pattern of linkage between variants in that population. Other populations have different patterns of linkage—including populations from the past. It is trivial to calculate a PRS score from the genotypes in an ancient sample. But whether that PRS score is meaningful depends on whether the population has been conservative enough in its linkage for the score to generalize. This means that PRS scores may be bad estimators of phenotypic differences between populations. A good review of the PRS appoach in ancient DNA is by Evan Irving-Pease and coworkers from 2021.
Akbari and collaborators looked at PRS scores for anatomical phenotypes like body fat percentage and waist circumference, mental health phenotypes like schizophrenia and bipolar disorder, and social phenotypes like smoking, household income, and years of schooling. In today’s societies, each of these is correlated weakly with genotypes across many genes, to a greater or lesser degree. But of course phenotypes like “years of schooling” and “current smoker” have no direct relevance to Bronze Age people. The aim of this part of the study was to assess whether the variants included within the PRS scores in samples today have changed in similar ways or diverse ways over time.
Akbari and coworkers found that the genotypes that comprise some PRS scores have evolved directionally across the last 10,000 years. The variants associated with smoking today have been declining in frequency across that time period. Variants associated with years of schooling today have been increasing in frequency.
What does this mean? Interpreting these polygenic results depends on how the trait and environment have changed. This post has already run pretty long. I’ll split the discussion of these polygenic cases of selection off into another post where I can take them in greater detail.
Moving forward
Studies of selection in humans across the last 10,000 years have often avoided grappling with the most interesting thing about humans. Humans are not passive actors responding to external changes in environment. Much of the pattern of recent selection happened within environments that people created themselves.
To be sure, there are textbook examples of coevolution that emphasize the cultural aspect of environments. There has been no selection for lactase persistence in cultures where domesticated dairy animals were not important to diet. Cultural innovation created the circumstances in which lactose digestion mattered beyond early childhood. Cultural transmission gave that dietary environment staying power across generations—even across thousands of years. Increased frequencies of new variants upregulating lactase were not the only outcome; an array of different technological and social innovations followed in different dairying societies.
But two aspects of coevolution are often missed.
One is that selection in a growing population is not zero-sum. After all, the measure of selection is the differential survival and reproduction of individuals who have a genotype. In a growing population, an increase in the frequency of a variant is the differential increase in numbers. People with dairy animals had more descendants at some times and places in part because some of them had lactase persistence variants.
The zero-sum expectation often leads to the misconception that selection diverges from population history. This misconception underlies some tests for selection, as researchers aim to find genes that violate the parameters of random genetic drift established by the population history. A better understanding is that selection was one of the causes of population history. Populations that grew often had many adaptive genetic variants contributing to their growth.
The other aspect of coevolution that is often missed is the social dimension. When people create environments, individuals have choices about how and whether to live in them, and with whom. The usual storytelling about selection ignores these choices. When the story of selection focuses on some new gene that helped conquer some environmental challenge, then it seems that people are passive victims. The mechanics of resistance to malaria, adaptation to high altitude hypoxia, synthesis of vitamin D, and digestion of lactose are all physical stories. But each of these also has social dimensions that help explain the presence and persistence of humans within challenging environments.
If we could look at all the people in the world 10,000 years ago, the people whose genes are most overrepresented today are those who chose to make larger social groups work despite the costs and tradeoffs, to tolerate diets centered on what previously were barely edible foods, and to care for, manage, and defend animals that started off wild and often resisted taming. Along the way, again and again their descendants faced unanticipated problems from these novel ways of life. They were subject to floods, pestilence, droughts, invading barbarians, the depredations of their own rulers, and the indifference of their gods. Many died. The barbarians often conquered. Still, again and again some of them invented their way out of those challenges. The land was often fat, and the consistent expansion of village or pastoral life into new spaces made it harder to persist along very different paths.
To be sure, adaptive changes to physical and molecular environments mattered. New variants of immune system genes, blood groups, metabolic and digestive processes, all helped people survive and have more surviving children. But the factors that mattered most were people choosing to strive within the societies that were growing, instead of walking away, and people allowing others to join their societies instead of excluding them. Whatever genotypes might have been slightly correlated with those choices over the centuries went along for the ride.
How many genes really mattered in these vast changes? In our work from 2007, we estimated that roughly one gene in seven might have been affected by positive selection in the last 40,000 years. Our rough estimate was for a long time the only one that tried to go beyond counting the limited number of variants that met a statistical threshold, to try to get at the whole picture. Still, the data were unsatisfactory in many ways. At that time we could say very little about how population history had mattered to linkage, which might have inflated our estimate. We had no ability to look at selection on standing variants at all, which depressed our estimate. I’ve never wavered in my thought that a large fraction of the genome was affected by positive selection in the recent past, but I couldn’t be sure whether we were within an order of magnitude of the whole effect.
So I was very surprised in the final version of the work by Akbari and coauthors to find they have a very similar estimate. They developed a good handle on the limitations of their method based on the more than 400 events they identified. Even with more than 15,000 genomes, the power of their method is limited. That means their observations are the tip of the iceberg of a larger number of variants that escape detection—either because selection was weaker, or selection varied over time and space, or because the variants were not yet common enough to register.
“Although the exact number of independent loci cannot be estimated with high confidence, we project that there are at least 3,800 independent signals of directional selection that are in linkage disequilibrium with the overwhelming majority of variants in the genome.”—Ali Akbari and coworkers
If they are in the right ballpark, the scope of selection in western Eurasia would roughly correspond to one-sixth of protein-coding genes.
Many questions remain to answer. One of the most important is the relation of selection not only to time but to space. In our 2007 work, we found that selection had been very important not only in Europe but also in East Asian and African populations. In fact we saw somewhat earlier strong selection in Africa than in the other regions, which we interpreted as the result of earlier population growth.
Ancient DNA evidence from East Asia has already confirmed a similar pattern. The same evidence from Africa will be challenging to attain, due to the weaker preservation of DNA over time at low latitudes. Nevertheless, as long as our picture of evolutionary change relies only on high-latitude groups, we will have a very incomplete idea of the real factors that changed our ancestors’ lives and biology.
Notes: I’ve seen many commentators complaining that Akbari and collaborators did not cite earlier work that is a foundation for their findings. I tend not to whine about that kind of thing. I write for everyone to read and learn, and my own research in this area certainly speaks for itself. Both Jonathan Pritchard and I had articles in Scientific American describing the foundational work on selection. My postdoctoral advisor Henry Harpending and Gregory Cochran summarized many of these findings in their 2009 book, The 10,000-Year Explosion.
I find that it’s valuable to link readers to a broader array of sources. Many of those sources raised issues that have not yet been completely resolved and are fruitful areas for further exploration. Some of this work got people pulling at threads that went in very different directions: into disease genetics, the genetic structure of polygenic traits, and the history of human populations. Naturally, in a blog post it’s inevitable that I’m leaving out a lot of people who contributed to the greater understanding of human adaptive evolution. But hopefully this helps to recognize the large number of researchers who have emphasized the recent rapid pace of natural selection in our species.
References
Akbari, A., Perry, A., Barton, A. R., Kariminejad, M., Gazal, S., Li, Z., Zeng, Y., Mittnik, A., Patterson, N., Mah, M., Zhou, X., Price, A. L., Lander, E. S., Pinhasi, R., Rohland, N., Mallick, S., & Reich, D. (2026). Ancient DNA reveals pervasive directional selection across West Eurasia. Nature, 1–6. https://doi.org/10.1038/s41586-026-10358-1
Aqil, A., Speidel, L., Pavlidis, P., & Gokcumen, O. (2023). Balancing selection on genomic deletion polymorphisms in humans. eLife, 12, e79111. https://doi.org/10.7554/eLife.79111
Barton, A. R., Rohland, N., Mallick, S., Pinhasi, R., Akbari, A., & Reich, D. (2026). Convergent natural selection at both ends of Eurasia during parallel radical lifestyle shifts in the last ten millennia. Evolutionary Biology. https://doi.org/10.64898/2026.04.03.716344
Bersaglieri, T., Sabeti, P. C., Patterson, N., Vanderploeg, T., Schaffner, S. F., Drake, J. A., Rhodes, M., Reich, D. E., & Hirschhorn, J. N. (2004). Genetic Signatures of Strong Recent Positive Selection at the Lactase Gene. The American Journal of Human Genetics, 74(6), 1111–1120. https://doi.org/10.1086/421051
Hawks, J. (2014). Still evolving (after all these years). Scientific American, (September), 86–91.
Hawks, J., Wang, E. T., Cochran, G. M., Harpending, H. C., & Moyzis, R. K. (2007). Recent acceleration of human adaptive evolution. Proceedings of the National Academy of Sciences, 104(52), 20753–20758. https://doi.org/10.1073/pnas.0707650104
Irving-Pease, E. K., Muktupavela, R., Dannemann, M., & Racimo, F. (2021). Quantitative Human Paleogenetics: What can Ancient DNA Tell us About Complex Trait Evolution? Frontiers in Genetics, 12, 703541. https://doi.org/10.3389/fgene.2021.703541
Mathieson, I., Lazaridis, I., Rohland, N., Mallick, S., Patterson, N., Roodenberg, S. A., Harney, E., Stewardson, K., Fernandes, D., Novak, M., Sirak, K., Gamba, C., Jones, E. R., Llamas, B., Dryomov, S., Pickrell, J., Arsuaga, J. L., de Castro, J. M. B., Carbonell, E., … Reich, D. (2015). Genome-wide patterns of selection in 230 ancient Eurasians. Nature, 528(7583), Article 7583. https://doi.org/10.1038/nature16152
Novembre, J., & Di Rienzo, A. (2009). Spatial patterns of variation due to natural selection in humans. Nature Reviews Genetics, 10(11), 745–755. https://doi.org/10.1038/nrg2632
Pritchard, J. K. (2010). How we are evolving. Scientific American, (October), 41–47.
Sabeti, P. C., Reich, D. E., Higgins, J. M., Levine, H. Z. P., Richter, D. J., Schaffner, S. F., Gabriel, S. B., Platko, J. V., Patterson, N. J., McDonald, G. J., Ackerman, H. C., Campbell, S. J., Altshuler, D., Cooper, R., Kwiatkowski, D., Ward, R., & Lander, E. S. (2002). Detecting recent positive selection in the human genome from haplotype structure. Nature, 419(6909), Article 6909. https://doi.org/10.1038/nature01140
Sams, A. J., Hawks, J., & Keinan, A. (2015). The utility of ancient human DNA for improving allele age estimates, with implications for demographic models and tests of natural selection. Journal of Human Evolution, 79, 64–72. https://doi.org/10.1016/j.jhevol.2014.10.009
Voight, B. F., Kudaravalli, S., Wen, X., & Pritchard, J. K. (2006). A Map of Recent Positive Selection in the Human Genome. PLOS Biology, 4(3), e72. https://doi.org/10.1371/journal.pbio.0040072
Wilde, S., Timpson, A., Kirsanow, K., Kaiser, E., Kayser, M., Unterländer, M., Hollfelder, N., Potekhina, I. D., Schier, W., Thomas, M. G., & Burger, J. (2014). Direct evidence for positive selection of skin, hair, and eye pigmentation in Europeans during the last 5,000 y. Proceedings of the National Academy of Sciences, 111(13), 4832–4837. https://doi.org/10.1073/pnas.1316513111